AlphaLink is a new workflow which uses AlphaFold2 with XL-MS for better protein structure predictions
10 min read
AlphaFold
Protein structures are defined, experimentally, by X-ray crystallography, nuclear magnetic resonance and electron cryomicroscopy (cryo-EM). However, these techniques are complex, time consuming, expensive and the protein is often not in its native form (Bertoline et al., 2023). Therefore, from its introduction in 2018, DeepMind’s AlphaFold program has quickly become an important tool in structural biological research to predict highly accurate protein structures based on their amino acid sequences (Bonislawski, 2023).
AlphaLink
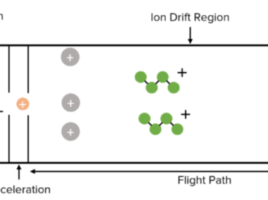
The latest version of AlphaFold, ‘AlphaFold2’, is typically used to predict single protein structures with high accuracy (Bertoline et al., 2023). A limitation of this computational technique is that predictions remain a challenge for proteins with little amino acid sequence data or those that undergo conformational changes (Stahl et al., 2023b). The first version of AlphaLink was proposed as a new workflow which incorporates data from cross-linking mass spectrometry (XL-MS), which provides information of the distance between specific amino acid residues, to aid AlphaFold2 in making better predictions (Stahl et al., 2023b). The study found that there was improved prediction accuracy of challenging single protein targets. However, predictions of protein complexes remained a challenge.
In 2022, DeepMind released ‘AlphaFold-Multimer’ which significantly increased the accuracy of protein complex prediction (Evans et al., 2022). This was quickly followed by the release of a new version of AlphaLink which used “AlphaFold-Multimer” rather than “AlphFold2”. The preprint paper showed again that prediction accuracy increased when XL-MS was incorporated into the workflow (Stahl et al., 2023a).
Integrating experimental data into computational algorithm development
With the increasing use of deep learning methods in analysing biological datasets, should data from systems such as XL-MS be incorporated into the development of neural networks in biological data analysis? AlphaLink uses XL-MS data to define restraint factors on the dataset which increases the accuracy of protein prediction by deep learning methods. Therefore, it is likely that when biological data is used during the design phase of machine learning algorithms more significant conclusions are made. But possibly the biggest limiting factor for setting up more interdisciplinary teams in biological research is the communication between experimental biologists and machine learning experts.
For more information:
- The impact of AlphaFold2 on protein prediction – https://www.ncbi.nlm.nih.gov/pmc/articles/PMC10011655/
References
BERTOLINE, L. M. F., LIMA, A. N., KRIEGER, J. E. & TEIXEIRA, S. K. 2023. Before and after AlphaFold2: An overview of protein structure prediction. Front Bioinform, 3, 1120370.
BONISLAWSKI, A. 2023. DeepMind’s AlphaFold Seeing Uptake for Protein-Protein Interaction Work [Online]. Available: https://www.genomeweb.com/proteomics-protein-research/deepminds-alphafold-seeing-uptake-protein-protein-interaction-work [Accessed 11 Aug 2023].
EVANS, R., O’NEILL, M., PRITZEL, A., ANTROPOVA, N., SENIOR, A., GREEN, T., ŽÍDEK, A., BATES, R., BLACKWELL, S., YIM, J., RONNEBERGER, O., BODENSTEIN, S., ZIELINSKI, M., BRIDGLAND, A., POTAPENKO, A., COWIE, A., TUNYASUVUNAKOOL, K., JAIN, R., CLANCY, E., KOHLI, P., JUMPER, J. & HASSABIS, D. 2022. Protein complex prediction with AlphaFold-Multimer. bioRxiv, 2021.10.04.463034.
STAHL, K., BROCK, O. & RAPPSILBER, J. 2023a. Modelling protein complexes with crosslinking mass spectrometry and deep learning. bioRxiv, 2023.06.07.544059.
STAHL, K., GRAZIADEI, A., DAU, T., BROCK, O. & RAPPSILBER, J. 2023b. Protein structure prediction with in-cell photo-crosslinking mass spectrometry and deep learning. Nat Biotechnol.